Evolutionary EcologyPollinators mediate microbiome assembly in flowers
In progress




Question
How pollinators mediate microbiome assembly in flowers is a major unresolved question of theoretical and applied importance in the face of anthropogenic disturbance and climate change.
Project Summary
Microbiomes can profoundly influence plant fitness in natural and agricultural settings. Relative to rhizosphere (root) and phyllosphere (leaf), the anthosphere (floral) microbiome is least well studied but has the most direct impact on plant reproductive success. Our understanding of what governs microbiome assembly in flowers – a highly dynamic niche – has advanced only recently. The identified drivers include, for instance, microbial dispersal mediated by pollinators, floral traits that can influence niche availability and microbial source pool, anthropogenic changes in mutualistic and antagonistic interactions, and disturbance imposed by agrochemicals. Yet, as these different drivers have often been studied independently, we lack a clear view as to how they act together and thus their relative importance in shaping the floral microbiome.
To address this question, we are using crop–wild relative systems in strawberries (Fragaria) and apples (Malus). This line of research involves a series of related projects: (1) What influences plant–pollinator interactions? (2) How do plant–pollinator interactions govern floral microbial diversity and network? (3) How do plant mutualistic (e.g. mycorrhizae) and antagonistic interactions (e.g. herbivory) at the rhizosphere and phyllosphere influence the floral microbiome? (4) How does phenology change influence plant–pollinator interactions and in turn the floral microbiome? We address these questions using field observations of pollinators, manipulative experiments (e.g. agrochemicals, mycorrhizae, herbivory), floral trait measurements (e.g. flower abundance, size, pollen production, color, volatiles), and microbiome sequencing.
Keywords
community assembly, crabapple, NCEP plot, microbiome, mycorrhizae, pollinators, phenology, strawberry
In the field or lab
Select Publications
- Fetters AM, Cantalupoa PG, Wei N, Roblesa MTS, Stephens JD, Pipasa JM and Ashman T-L. 2022. The pollen virome of wild plants and its association with variation in floral traits and land use. Nature Communications.
- Wei N, Whyle RL, Ashman T-L and Jamieson MA. 2022. Genotypic variation in floral volatiles influences floral microbiome more strongly than interactions with herbivores and mycorrhizae in strawberries. Horticulture Research.
- Wei N, Kaczorowski RL, Arceo-Gómez G, O’Neill EM, Hayes RA, Ashman T-L. 2021. Pollinators contribute to the maintenance of flowering plant diversity. Nature 597:688-692.
- Wei N, Whyle RL, Ashman T-L and Jamieson MA. Genotypic variation in floral volatiles influences floral microbiome more strongly than interactions with herbivores and mycorrhizae in strawberries. Horticulture Research. In Review.
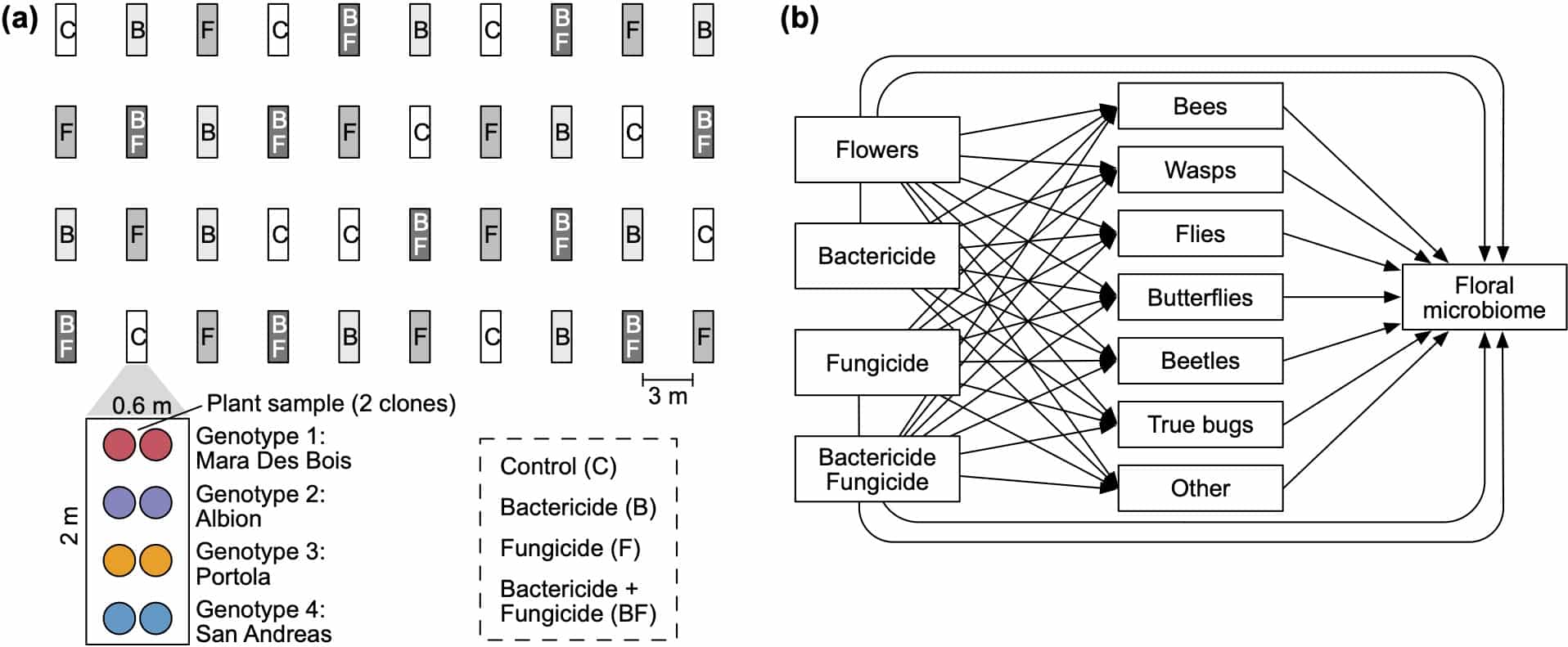
- Wei N, Russell AL, Jarrett AR and Ashman T-L. 2021. Pollinators mediate floral microbial diversity and network under agrochemical disturbance. Molecular Ecology 30:2235-2247.
- Cullen N, Xia J, Wei N, Kaczorowski RL, Arceo-Gómez G, O’Neill EM, Hayes RA and Ashman T-L. 2021. Diversity and composition of pollen loads carried by pollinators are primarily driven by insect traits, not floral community characteristics. Oecologia 196:131-143.
- Wei N, Incarnato M and Kaufman E. 2021. Plant–pollinator interactions in crabapples: A case study at Holden Arboretum and Secrest Arboretum, Ohio. Malus: International Ornamental Crabapple Society Bulletin.
- Wei N, Kaczorowski RL, Arceo-Gómez G, O’Neill EM, Hayes RA and Ashman T-L. 2020. Pollinator niche partitioning and asymmetric facilitation contribute to the maintenance of diversity. bioRxiv. https://doi.org/2020.03.02.974022.
- Rebolleda-Gómez M, Forrester NJ, Russell AL, Wei N, Fetters AM, Stephens JD and Ashman TL. 2019. Gazing into the anthosphere: considering how microbes influence floral evolution. New Phytologist 224:1012-1020.
- Wei N and Ashman T-L. 2018. The effects of host species and sexual dimorphism differ among root, leaf and flower microbiomes of wild strawberries in situ. Scientific Reports 8:5195.











